FLUNAMI
Calling for more public health research

UQ Child Health Research Centre Virologist Associate Professor Ian Mackay and Honorary Lecturer Dr Katherine Arden discuss the unseasonal flu wreaking havoc on emergency departments across Australia and the changing tide of these evolving viruses.
Australians have been pummelled by a rising tide of flu cases since late 2018 and science has yet to produce any answers to a raft of questions.
Flu viruses are continually mutating. They divide their protein-making genetic blueprint into eight parts. This format conveys an ability to swap out whole genes, potentially changing viral appearance and behaviour more quickly than by mutational drift alone. There are two types of flu virus that cause regular seasonal human epidemics; A and B. FluA strains are subtyped; A/H3N2 – the most diverse today – and A/H1N1. FluB strains exist in lineages; B/Victoria and B/Yamagata.
Rising tides in recent years
Around most of Australia, 2017 was the biggest year of lab-confirmed flu ever recorded. A/H3N2 dominated, but FluB strains also co-circulated, eventually commanding the end of the season. We now know that the 2017 A/H3N2 component of the vaccine mutated during production so the 2017 vaccine didn’t offer the protection it could have. On top of that, wild viruses evolved in a different direction.
Over 1,200 deaths were recorded, mostly due to FluA among the elderly. This heavy toll is why stronger vaccines now exist. These high dose vaccines help prevent severe consequences of infection among those 65 years or older. Despite high numbers, the rate of death didn’t differ from what is seen every year.
In 2018, flu took a break. Activity and impact were low, and severity was moderate.
Then came the summer of flu...
From late 2018, we saw a rare occurrence for flu; it reached record levels during summer. The last national rise like this was in 1995, but Australia has never recorded so many confirmed flu cases as we have in in the first half of 2019. This year, June alone recorded more cases than any previous year’s peak month, except for 2017, and May recorded nine times more cases than the next highest May on record.
Hospitals have been strained, not because of exceptionally severe flu cases, but due to the sheer number of cases at an unexpected time. In most Australian jurisdictions, flu has been circulating above the usual baseline for more than seven months.
"Hospitals have been strained, not because of exceptionally severe flu cases, but due to the sheer number of cases at an unexpected time."
Growing evidence suggests the largest group of A/H3N2 strains currently in Australia may not be an ideal match for the A/H3N2 used as the southern hemisphere vaccine component; A/H3N2 may have outsmarted our vaccine selection and production process again.
There has been no research to explain the early flu storm
Infected travellers carry new flu strains, but they always have, and tourist visits and returning traveller numbers haven’t spiked over summer.
Testing has become easier with rapid point-of-care methods added to hospitals in recent years. We do more lab testing of samples now than we did before 2001, when flu became a notifiable disease. But remember 2018? It proved that a small flu year was still a small flu year, regardless of test availability.
Could extreme temperatures have crowded us indoors, increasing transmission? Perhaps, but we’ve had frequent hot summers lately.
For flu numbers to be high so early, something has changed. The key pieces of the puzzle left unstudied are the viruses. We have very little virological detail about them.



H1N1 virus
We need more public health research to modernise flu characterisation.
Flu is different every year, yet we agonise, guess and listen to others guess about the trajectory of a given season. For a concerned public, there are often gaping communication holes, devoid of science-based commentary and analysis but quickly flooded with sensationalism.
Summer 2018/19 flu activity raises questions:
- Was anything different about the flu viruses circulating when the first wave began?
- Are A/H3N2 viruses diversifying more rapidly than ever?
- Could we identify the start of a pandemic early?
- Are we taking seriously the importance of public communication?
Whole Genome Sequencing could hold the answer to these new strains of flu virus.
We could address these questions if we modernised our approaches. In particular, whole genome sequencing (WGS) – no longer a young technology – rapidly and reliably describes all viral genes at once. WGS could allow scientists to better quantify co-circulating flu viruses and identify new recombinants; viruses that have replaced one or more of their genes for those of a co-infecting flu virus. More extensive data could characterise an epidemic and its likely impact on public health measures. WGS and expertise in its interpretation would inform vaccine formulations, strengthen understanding of worldwide flu movements, identify and prepare us for future pandemics and generally improve our flu virus understanding.
This year it was Flunami, next year a different flu story. Any number of potentially pathogenic viruses could also emerge, arrive or reach epidemic proportions in the meantime. Public health research needs to grow and respond more flexibly to viral threats. To do so, it must emulate the flexibility of its viral targets, or it may fail to protect and inform the public it serves.

H1N1 virus
H1N1 virus
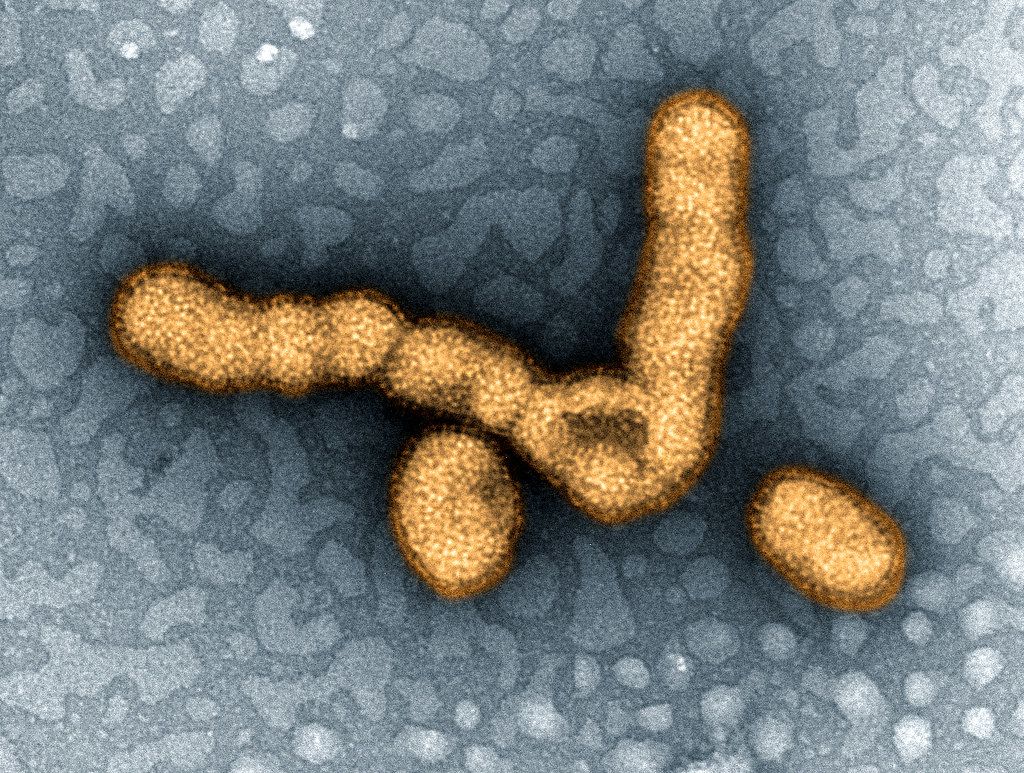
Understanding the virology is key to providing better management and treatment of the flu.
Understanding the virology is key to providing better management and treatment of the flu.

Whole Genome Sequencing could hold the answer to these new strains of flu virus.
Whole Genome Sequencing could hold the answer to these new strains of flu virus.
